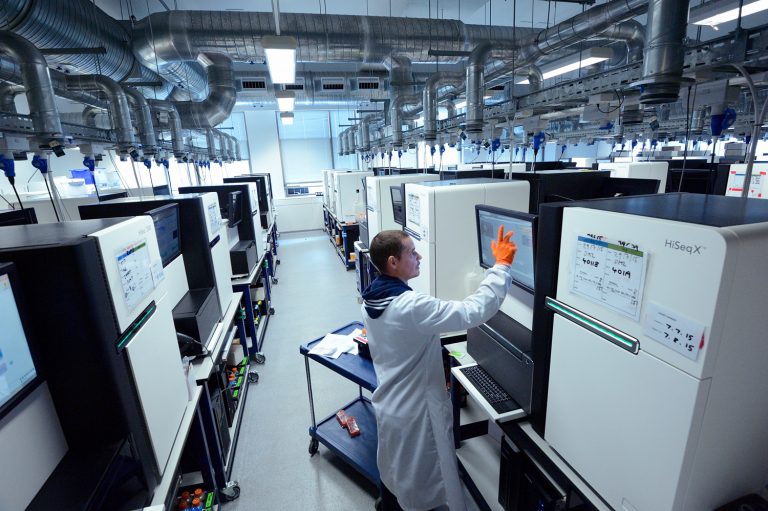
Nanorate sequencing, developed at the Wellcome Sanger Institute in the U.K., is extremely accurate and allows detection of new mutations in non-dividing cells, something that was difficult with earlier techniques.
The sequencing method, also known as NanoSeq, uses duplex sequencing. This involves random molecular tagging of double-stranded DNA to improve accuracy and allow better detection of mutations.
Duplex sequencing is not new, but earlier iterations of the technique made it very difficult to detect mutations only present in single cells or small sections of non-dividing tissue. Mutations occur over time in our non-germline cells (at a rate of 15-40 per year) and these are often only in single or small groups of cells. While many such variants are harmless, some may cause cancer or other diseases later on.
This new NanoSeq technique took 4 years to develop and involved a careful refining of currently used duplex sequencing techniques. For example, using more specific enzymes to cut DNA more accurately and also better bioinformatics analysis of the sequencing data. It has an error rate of less than 5 errors per billion base pairs in single DNA molecules from cell populations. It also does not require invasive tissue biopsies and samples can be taken using skin or tissue swabs.
Robert Osborne, Ph.D., previously based at the Sanger, and Inigo Martincorena, Ph.D., a group leader at the Sanger led the research, which is published in Nature.
“Detecting somatic mutations that are only present in one or a few cells is incredibly technically challenging,” said Osbourne, in a press statement.
“You have to find a single letter change among tens of millions of DNA letters and previous sequencing methods were simply not accurate enough. Because NanoSeq makes only a few errors per billion DNA letters, we are now able to accurately study somatic mutations in any tissue.”
The team tested their method to evaluate levels of mutations in stem cells and non-dividing cells in the blood, neurons and colon.
They found similar levels of mutations in adult, differentiated blood and colon cells compared with stem cells in the same tissues. This was surprising for blood cells as they undergo frequent cell division. Neuronal cells, in contrast, seemed to gradually accumulate mutations over time.
“It is often assumed that cell division is the main factor in the occurrence of somatic mutations, with a greater number of divisions creating a greater number of mutations. But our analysis found that blood cells that had divided many times more than others featured the same rates and patterns of mutation. This changes how we think about mutagenesis and suggests that other biological mechanisms besides cell division are key,” said Federico Abascal, Ph.D., a researcher in Martincorena’s group at the Sanger and the first author of the paper.
The researchers are now planning to develop and refine the technique further in more types of tissue and cell samples. “The application of NanoSeq on a small scale in this study has already led us to reconsider what we thought we knew about mutagenesis, which is exciting,” says Martincorena.
“NanoSeq will also make it easier, cheaper and less invasive to study somatic mutation on a much larger scale. Rather than analysing biopsies from small numbers of patients and only being able to look at stem cells or tumour tissue, now we can study samples from hundreds of patients and observe somatic mutations in any tissue.”